The hypothesis-testing paradigm
We all remember our first science fair project. All you wanted to do was set fire to something or make it explode. But before you were even allowed to touch the baking soda and vinegar, your teacher made you come up with a clear hypothesis that you were going to test - because carrying out an experiment without a specific question in mind would be pointless right? Well, maybe not.
While this particular method of ‘hypothesis-testing’ based inquiry has been an essential part of many scientific discoveries, rapid advancements in technology has opened up a new realm of hypothesis-generating approaches, where high-throughput screening methods such as genomics and proteomics allow you to generate enormous amounts of data in the space of just a few days. In this way, scientists can ask unbiased questions by letting the biology drive their hypotheses.

Research Fellow Dr. Melanie White from the University of Sydney has been integrating these hypothesis-generating approaches into her research for the last 20 years. In particular, her lab combines the time-tested Langendorff perfusion technique with proteomics, to understand how the heart adapts to altered physiological states such as hyperglycemia in a diabetic heart.
Melanie explains that ‘With diabetes being a global epidemic and its inherent associated risk with heart disease - it’s a burning biological question. In the clinic, patients with diabetes are at a greater risk of death following myocardial ischemia. So, our primary question is, what happens in the diabetic heart to prime this failure to recover?'
'Currently, we are investigating changes in signaling cascades in diabetic hearts. We know that metabolic pathways are altered but we suspect that there are deeper adaptations at play, ones that are yet to be appreciated because of the way in which hypotheses are developed and then tested. We use proteomics as a hypothesis-generating approach, and in this way, the biological data drives our hypotheses and follow up targets.’
The power of proteomics
Proteomics is like taking a polaroid picture, a snapshot of all the proteins present in a cellular environment, at a particular point in time. By quantifying the relative abundance, or presence/absence of various proteins under different conditions i.e. in a normal healthy heart vs a diabetic heart, you can begin to understand what changes are happening within the cells to cause this pathological state.
Identifying these differences can be extremely useful in the development of new drug targets for the treatment of disease, or in the creation of better diagnostic tests that are highly specific to certain pathologies.
‘For our proteomic work, we take intact proteins and break them into their constituents, or peptides, using controlled proteolysis. Beyond this, we analyze these peptides based on their intact mass and the masses of the amino acids that comprise them, using high-resolution, high-sensitivity mass spectrometry - which can make “mass maps”. Tens of thousands of precise mass measurements in the space of an hour or two.’
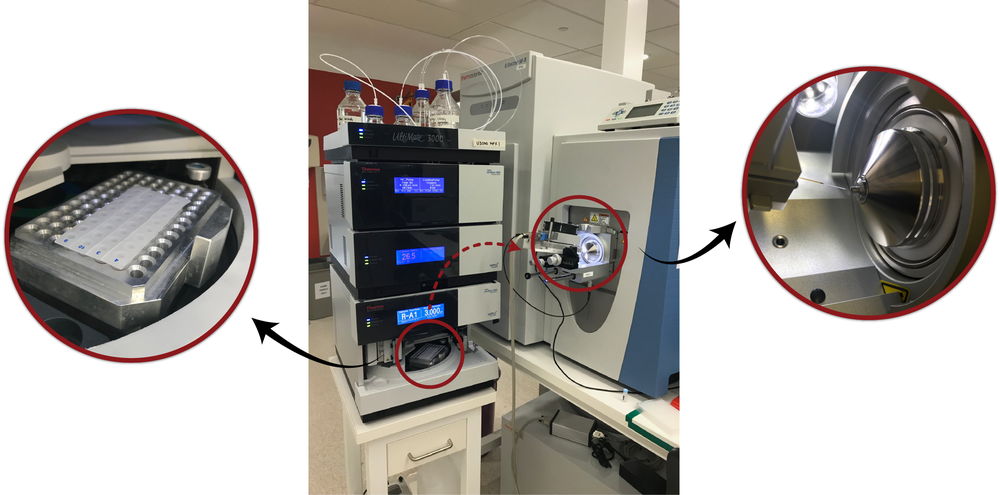

The White Lab’s Liquid Chromatography-Mass Spectrometry (LC-MS/MS) System (center). Peptide samples obtained from the heart tissue are placed in an insulated holder (left) where they are subjected to high-pressure liquid chromatography for separation. The samples are then ionized for injection into the mass spectrometer (right).
‘We then compare these mass maps to in-silico predictions of the peptides that arise from a translated genome. It is the developments in this field that allow us to drill down into the cell and ask how signal cascades are regulated, how well the cell copes with redox stress, how proteins are broken down, or which proteins interact with each other. The list of possible questions goes on and on!’
Preparing the tissue for proteomics using Langendorff perfusion
Melanie and her team use isolated Langendorff heart perfusion to prepare their tissue samples for proteomic analysis. This technique is a crucial part of their experimentation, as it allows them to closely monitor the heart up until the sample is taken, as well as remove any obfuscating proteins (such as albumin in the blood) that would interfere with the proteomic analysis.
‘The ADInstruments technology is instrumental to our work through the generation of all our tissue samples. The system also allows us to collect extruded perfusate which has come into direct contact with cardiomyocytes, providing a novel way of investigating cardiac-specific biomarkers. Using the Langendorff system, we can precisely time our injury or the delivery of a therapeutic and closely observe the functional effects via digital recordings using PowerLab and LabChart. It also provides our checks and balances to support our proteomics work.’
Related: An Introduction to Isolated Langendorff Heart Perfusion
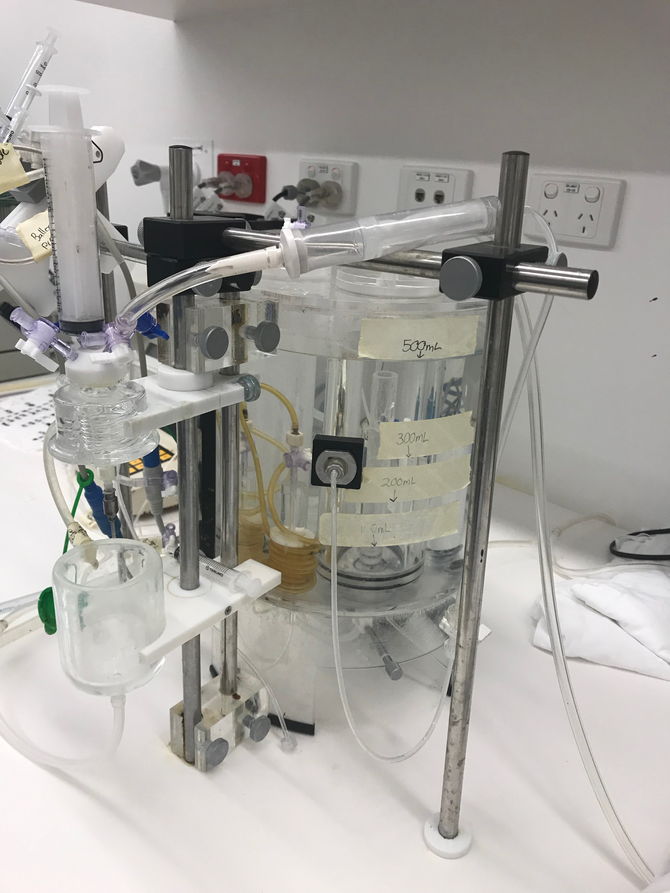
The White Lab's Panlab Langendorff Setup
During this period on the Langendorff Apparatus, they monitor physiological parameters such as perfusion pressure, flow rate, heart rate, diastolic and systolic pressure as well as calculating the Rate Pressure Product (RPP) using LabChart.
Read more: LabChart for Isolated Perfused Hearts.
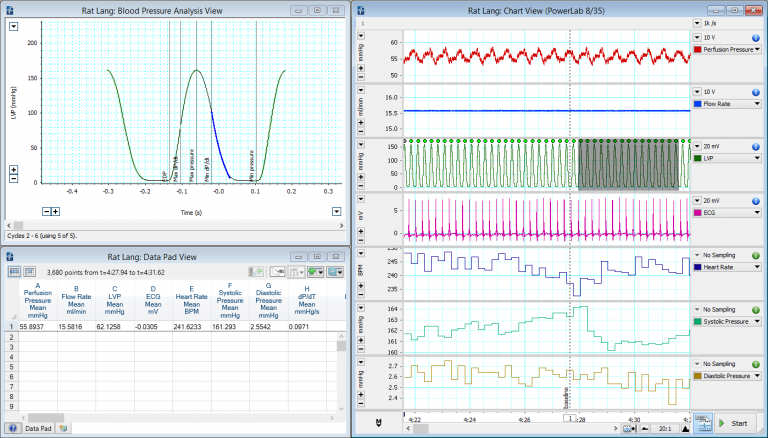
A typical Langendorff recording highlighting LVP, ECG, perfusion pressure and calculated variables.
'Proteomics paints a very clear picture of the cellular condition at a given moment in time. For this reason, our exclusion criteria are essential, and this is also where data acquired via LabChart is vital to our success. Using our proteomic approaches, we have seen the effects of “new hands” in the surgical excision of the heart, and the effects of arrhythmias just prior to the initiation / completion of a perfusion protocol. That’s the thing with proteomics - it will tell you something’s wrong. The power to go back and re-interrogate that initial data is so important to everything we do downstream.’
A key advancement in this area has been in the digitization of the Langendorff system. When Melanie first started out in the field, all the physiological parameters mentioned above we recorded using a Grass-Instrument Co. polygraph - generating reams and reams of paper that were exceedingly difficult to keep track of and manage! Now it’s just a case of hitting the save button.
‘I have been relying on Langendorff perfusion of hearts for nearly 20 years and have seen the field move rapidly in the last 10-15 years, mainly through the digitalization of data acquisition and the “miniaturization” of the setup. Having a desktop solution for Langendorff perfusion has revolutionized the field.’
The busy life of a researcher
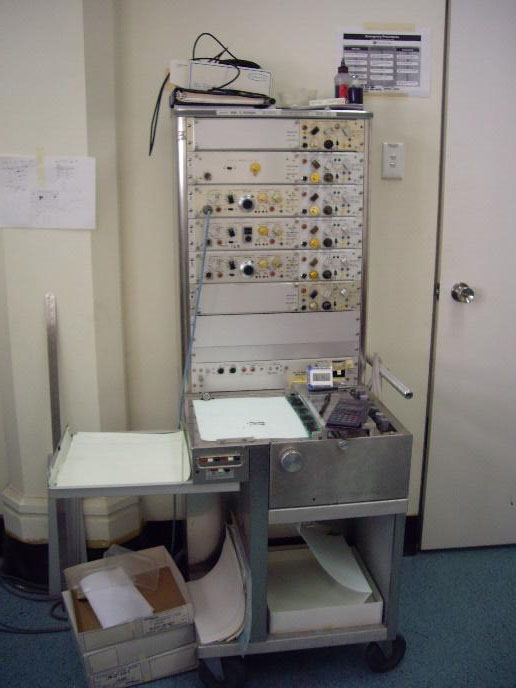

The old Grass-Instrument Co. polygraph
Building a successful career as a researcher is far from easy, it can take a lot of dedication and hard work. Like most primary investigators, Melanie’s time is often taken up by lecturing, applying for grants or supervising her students, however, she still finds the time to do some of the things she loves most.
'I still love going into the Lab (when they’ll let me!). I try to get in there as much as possible, as it keeps my curiosity burning. The other thing I really love doing is talking to people about my work. Sometimes just having a water cooler conversation can prompt you to go back and look at your data with a fresh perspective.'
When asked what she was most proud of, from her career so far, Melanie responded -'This may sound rather non-scientific, but I’m proud to be a working mum. Often having children and an active science career can be tricky to coordinate, and you need a really good support network around you. So being able to come back and do what I love is really rewarding.’
Dr. Melanie White is a Heart Foundation (Australia) Future Leader Fellow in Charles Perkins Centre, at the University of Sydney. She teaches senior students in the discipline of Pathology in the Faculties of Science and Medicine & Health.
Related:
Learn more about our complete Langendorff perfusion systems
Choosing the right perfusate for your isolated Langendorff heart preparation
Langendorff vs. Working Heart Perfusion – What’s the difference?
Effects of constant flow vs. constant pressure perfusion on Langendorff isolated heart studies


